This spring I've been fortunate to advise two student groups in computer science at UiT The Arctic University of Norway in Bodø. There's already a news post in Norwegian about this year's students on the DIPS website (in Norwegian), but I wanted to go into a bit more detail on what they worked on.
Last year I had a group of four students, who worked on machine learning with electronic health record data at DIPS. Unfortunately I did not take my time to summarize their work, and that's something I want to fix this year. If you want to read a short summary of what the students did last year, you can read the news post on the DIPS website (in Norwegian as well).
The students worked on pretty different problems. The first team was given the challenge to develop a system that allows Withings users to share their electrocardiograms (ECGs) with our patient journal system. The second team was given the challenge to extract relevant clinical information from unstructured clinical texts.
The two groups started their work this January, and delivered their bachelor theses a couple of weeks ago. I met with them every week to catch up on the development, give some advise, and help them solve their tasks. I'm really impressed by the work that the students put in, and the results that I present in this short blog post are their work. Here are the two teams that I was fortunate to advise:
Team ECG: Johanne Holst Klæboe, Sander Holdal, Ole Jørgen Hongset, Kristoffer Jakobsen
Team NLP: Christopher Hansen Gabrielsen, Kristoffer S. Dahl, Ådne Wolden Ovesen, William Harald Elvis Aamodt
Structured ECGs
The goal of this project was to store ECGs from Withings smart watches into our openEHR server at DIPS. Traditionally ECGs have been stored as static PDFs in the electronic health record systems, but in this project we wanted to explore the possibility of storing ECGs as structured data. Storing the raw ECGs makes it possible to re-use the data later, and significantly reduces the storage requirements needed to store an ECG.
In DIPS we store structured clinical data as openEHR archetypes, so a part of the student's work is to transform the ECGs from Withings' API into openEHR archetypes. If you want an intro to what openEHR is, I recommend that you read John Meredith's post on the topic. In short, it's how we model clinical concepts, and describes how we can e.g. store the results from an ECG.
The project consisted of five main parts;
- authorize an app to get a person's ECGs,
- fetch ECGs from the Withings API,
- convert them to openEHR archetype data,
- store them in the openEHR server
- develop a simple visualization dashboard that fetches the stored ECGs from the openEHR server, and visualizes the ECGs.
Withings provide raw ECG measurements from their API. The students set up an application to fetch data, and the necessary OAuth flow to let user's authorize the app to get their ECGs. Once retrieved, the students had to convert the raw measurements into a data format based on openEHR archetypes. With help from Bjørn Næss we landed on an initial approach to represent the ECGs. In short the data we're storing in our openEHR server is a Result report with the ECG result, along with information about the device that was used to collect the ECG. Data data is serialized as JSON and posted to our openEHR server via the openEHR REST API
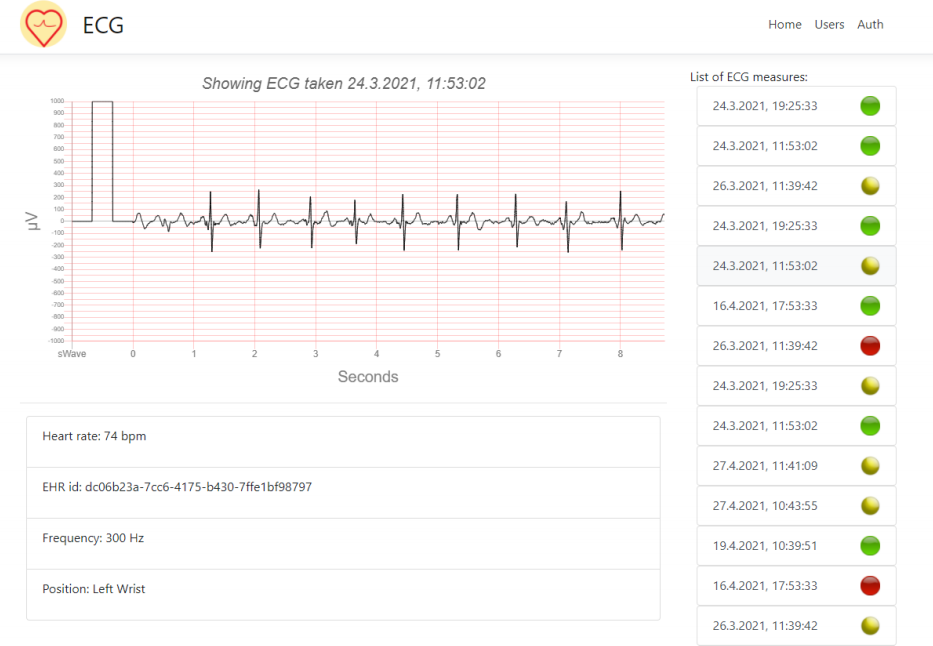
When the data is stored in our openEHR backend, the students used the Archetype query language (AQL) to retrieve all ECGs to visualize them in the web user interface. Below is an example from the web application that was developed by the students. It shows an interactive ECG visualization, with the possibility of navigating between different ECGs recorded by this patient.
Unstructured clinical text
In electronic health record systems, a lot of the information is stored as unstructured free text documents and notes. While this is great for using a clinician's natural language, it becomes difficult to find and extract specific parts of the document that may be of interest. E.g. finding blood pressures, symptoms or medications in the free text.
In this project the students were given the challenge to discover and extract important bits of information from unstructured clinical text. While real clinical texts may be difficult to get access to, my colleague Pål Brekke got access to ~30 000 articles from The Journal of the Norwegian Medical Association. These articles contain case studies with descriptions of patient's disease, and we hypothesized that these are similar to text found in the journal.
The students were given the NorMedTerm lexical resource resource of medical entities mapped to semantic categories. Based on these terms, they set out to develop a system that automatically extracted these terms from the text, and presented them to the user.
In short, the project consisted of four main parts:
- fetch the NorMedTerm lexical resource for later lookup,
- parse the ~30 000 articles stored as XML files,
- discover the relevant entities in the articles based on NorMedTerm
- present the articles and discovered entities to the user in a user interface
NorMedTerm is available as a .csv on their Github page, and is distributed
under a CC BY 4.0 license. Based on NorMedTerm, the students developed an API
that takes a clinical note, finds all medical terms, and returns a description
of the entities and their positions in the text. Based on this API, they
developed an interactive web application that allows users to browse the ~30 000
articles, and the discovered entities on the same page.
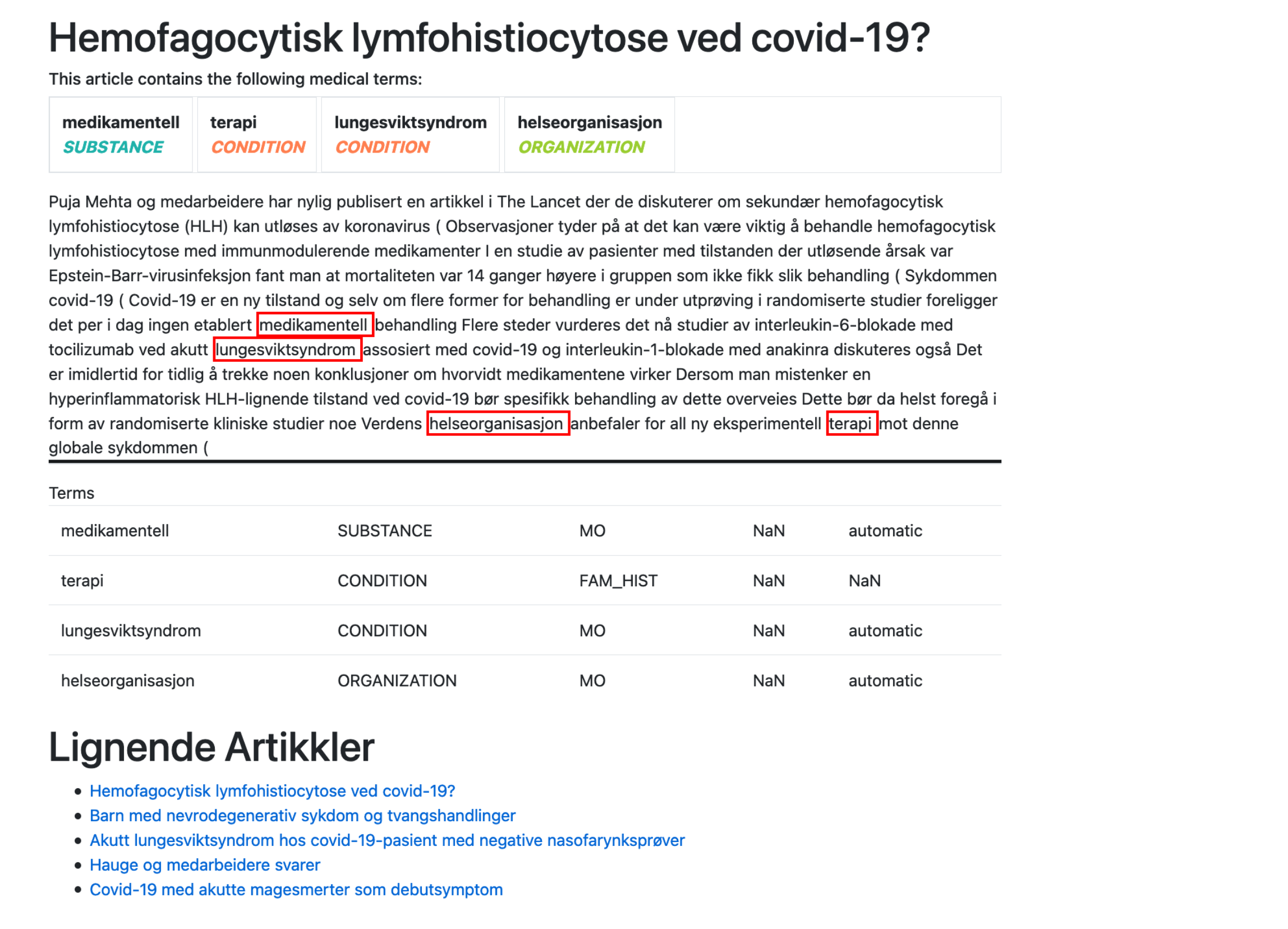
Below is a screenshot from the web application. The user is browsing an article on Covid-19, and relevant substances, conditions and other entities are highlighted in the text. Based on these findings, the website could also hint to other similar articles.
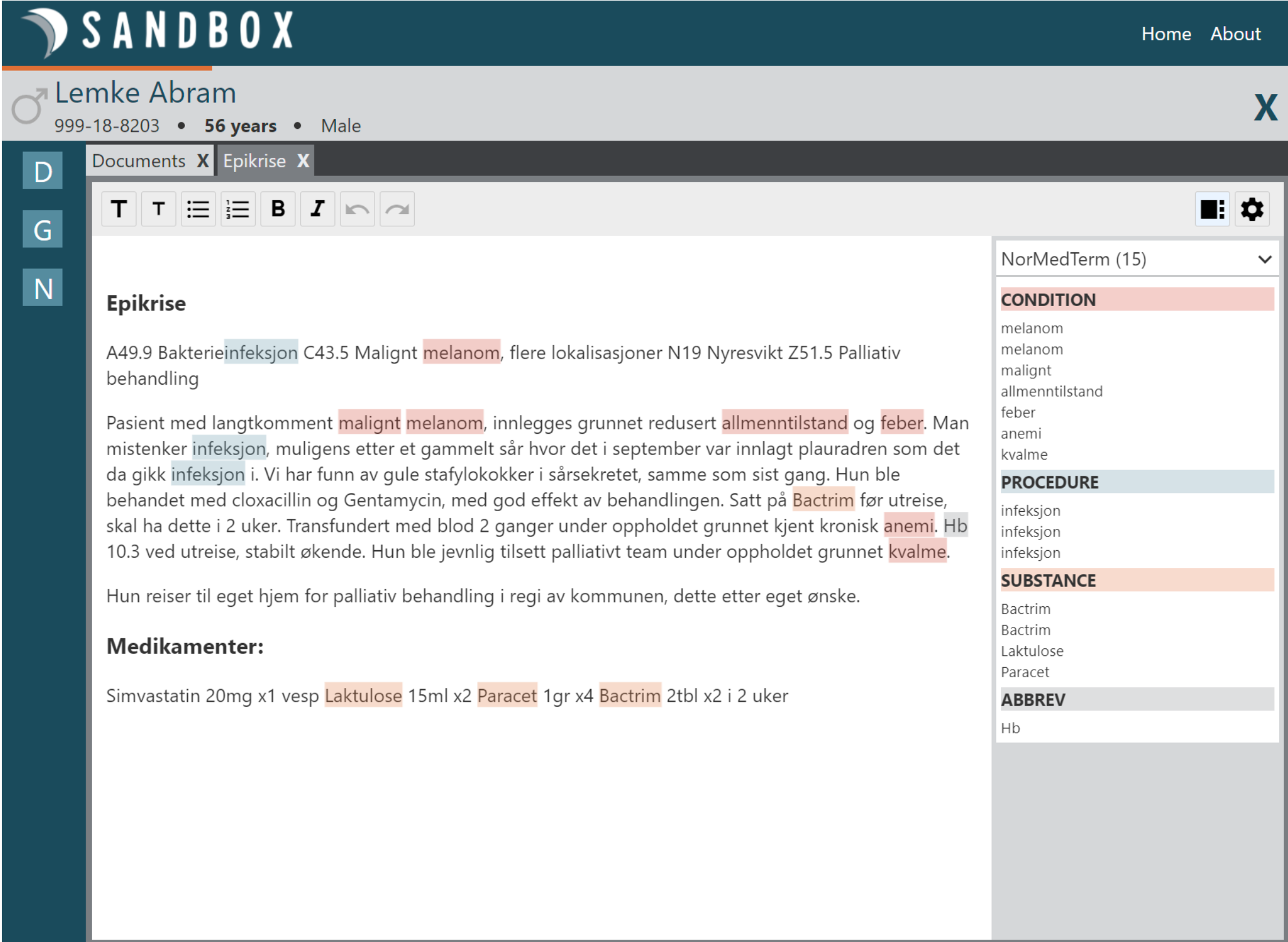
Since the students developed an API for discovering entities, my colleague Thor Stenbæk used their API in his new smart text editor in DIPS. Below you'll see a screenshot of how that looks like. Pretty cool!
Summary
In summary, the two groups have worked on different topics and built some pretty cool stuff. I believe that the structured ECGs are just the beginning of how we can start integrating personal data into the electronic health record system. Using natural language processing techniques on clinical notes is definitely something we're going to see more in the future.
Hopefully I'll get my hands on more students to do cool stuff next year as well!